background | research aims | techniques | infrastructure | useful links
background
Fibroblast heterogeneity in skin homeostasis
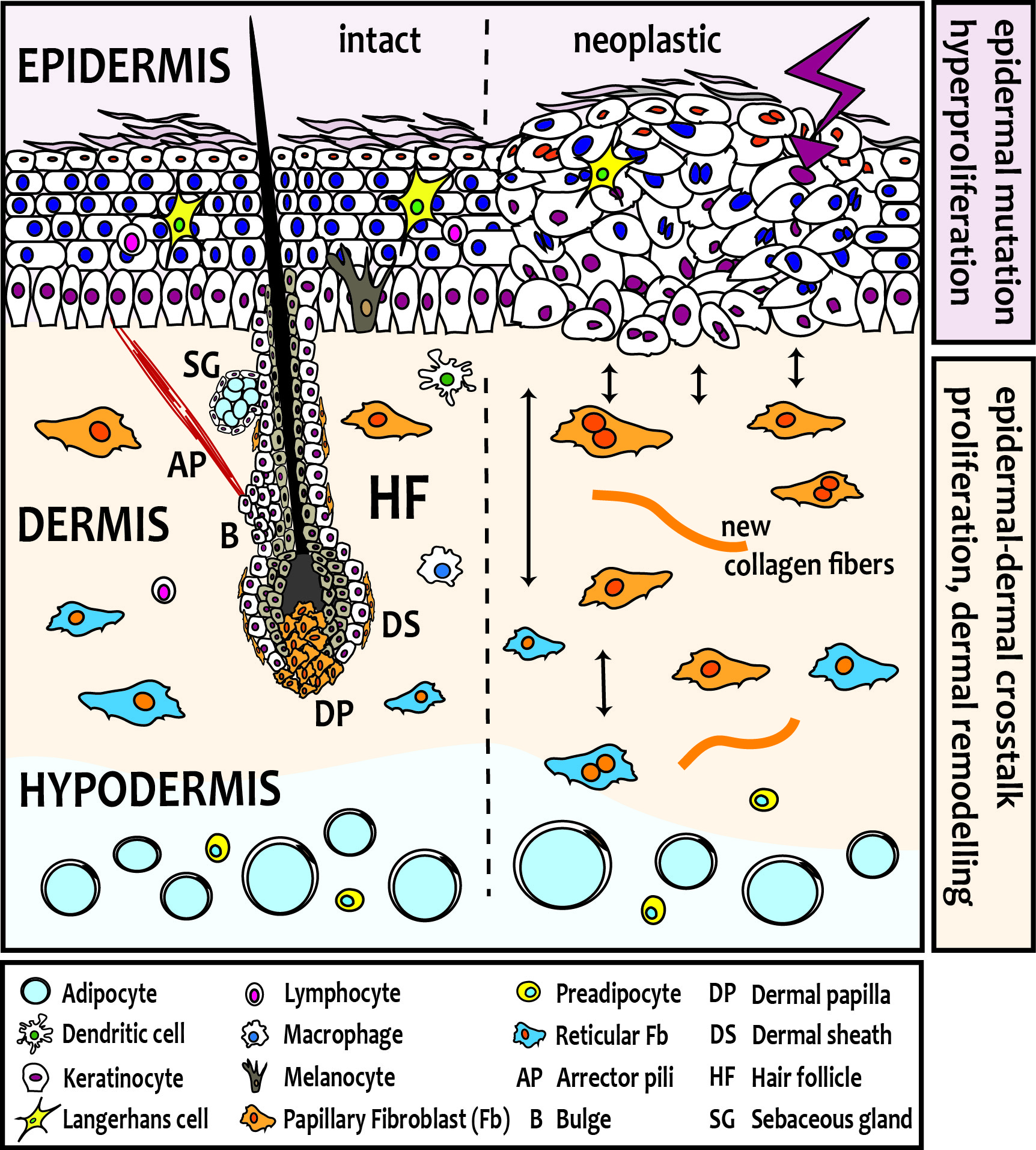
Skin fibroblasts arise from two distinct lineages during embryogenesis, which have unique functions in skin development and homeostasis. In mouse skin, one lineage gives rise to the upper, papillary dermis, including fibroblasts of the dermal papilla, dermal sheath and arrector pili muscle, required for hair follicle (HF) formation. The other lineage forms the lower dermis, including the reticular fibroblasts that synthesize the bulk of the fibrillar extra-cellular matrix (ECM), and the preadipocytes and adipocytes of the hypodermis. Several lines of evidence suggest that also human skin comprises at least two fibroblast lineages with distinct functions.
The papillary and reticular fibroblast lineages also play different roles in skin regeneration. While reticular fibroblasts representing the majority of fibroblasts in adult tissue are important for the first phase of wound healing, the papillary lineage is not repopulated until re-epithelialization and contributes exclusively to the upper dermis, which explains the absence of hair follicles in newly closed wounds and scarring. As skin ages, the papillary fibroblasts diminish in number leading to a progressive thinning of the skin, and besides the gene signature of fibroblasts changes, and thus their properties.
research aims
Functional deconstruction of distinct fibroblast subsets in skin physiology and disease
When skin tumours arise from neoplastic epidermal cells, they elicit profound and distinct changes in the dermis. Our studies are shedding light on the important issue of how signature oncogenic mutations in epithelial cells reprogram fibroblasts to cancer-associated fibroblasts (CAFs) and result in characteristic stromal responses, and if distinct fibroblast subsets have unique functions in skin cancer (basal and squamous cell carcinoma, melanoma) development and progression.
In addition, our lab is also investigating if modulating dermal signalling can redirect regeneration upon injury to a more scarless phenotype. Furthermore, we aim to address the role of the distinct fibroblast subsets in fibroblast-mediated skin pathologies such as scleroderma and keloids, which lack an effective clinical treatment regimen.
techniques
We use a broad range of methods including state-of-the-art cell and molecular biology methods, 3D in vitro culture systems (human skin equivalents, organoid cultures), next generation sequencing, single-cell transcriptomics, flow and mass cytometry (CyTOF), in vivo lineage tracing techniques and cutting-edge imaging technologies, and conditional knock-out mice or murine skin disease models to unravel the role of distinct fibroblast subsets in skin physiology and pathology.
infrastructure
Access to patient material
The Skin and Endothelium Research Division (SERD), which is part of the Department of Dermatology at the Medical University of Vienna (MUV), is located at the Anna Spiegel Center of Translational Research (ASCTR). The research groups at the SERD and the Department of Dermatology perform both basic research as well as translational studies mostly in collaboration with clinicians at the general hospital and the MUV, which is located at close proximity.
Core Facilities
Besides basic equipment for molecular and cellular biology studies, scientists at the ASCTR have access to all cutting-edge methods in biomedical research including state-of-the-art core facilities for histology, imaging, flow cytometry, genomics and proteomics. Furthermore, an SPF animal facility is available housing transgenic, knock-out and immune-deficient mice.
useful links
http://www.hair-gel.net/ Hair-GEL
http://www.sanger.ac.uk/science/collaboration/mouse-resource-portal/ Sanger Mouse Portal
http://www.mousephenotype.org/data/order/creline/ CREATE Cre lines
http://www.mousephenotype.org/ International Mouse Phenotyping Consortium
http://www.hipsci.org/ Human Induced Pluripotent Stem Cell Initiative
http://www.spotthedot.org/ Spot the Dot Foundation
http://www.bdbiosciences.com/br/research/multicolor/spectrum_viewer/ BD Fluorescence Spectrum Viewer for Multicolor Flow Cytometry
http://www.cancergenome.nih.gov/ The Cancer Genome ATLAS